I am an AI for Science researcher at the Lotfollahi Lab, Wellcome Sanger Institute, working at the intersection of machine learning and biomedical science.
My background in bioinformatics and biochemistry shapes how I approach this space — not by fitting biological data into existing AI frameworks, but by asking what a problem actually requires before reaching for a method. I focus on translating the complexity of biological and medical questions into well-defined computational problems, and building the models to solve them.
“What I cannot create, I do not understand”
- Richard Feynman
Academic CV
Current research interest
•
Multi-modal learning of spatial gene expression and histopathology image
•
Uncertainty-controlled de novo regulatory sequence design
Work experience
•
Senior Data Scientist at Sanger Institute
•
Education
•
•
◦
Selected as Fellow of the National Excellence Scholarship (Natural Sciences and Engineering)
Publication
•
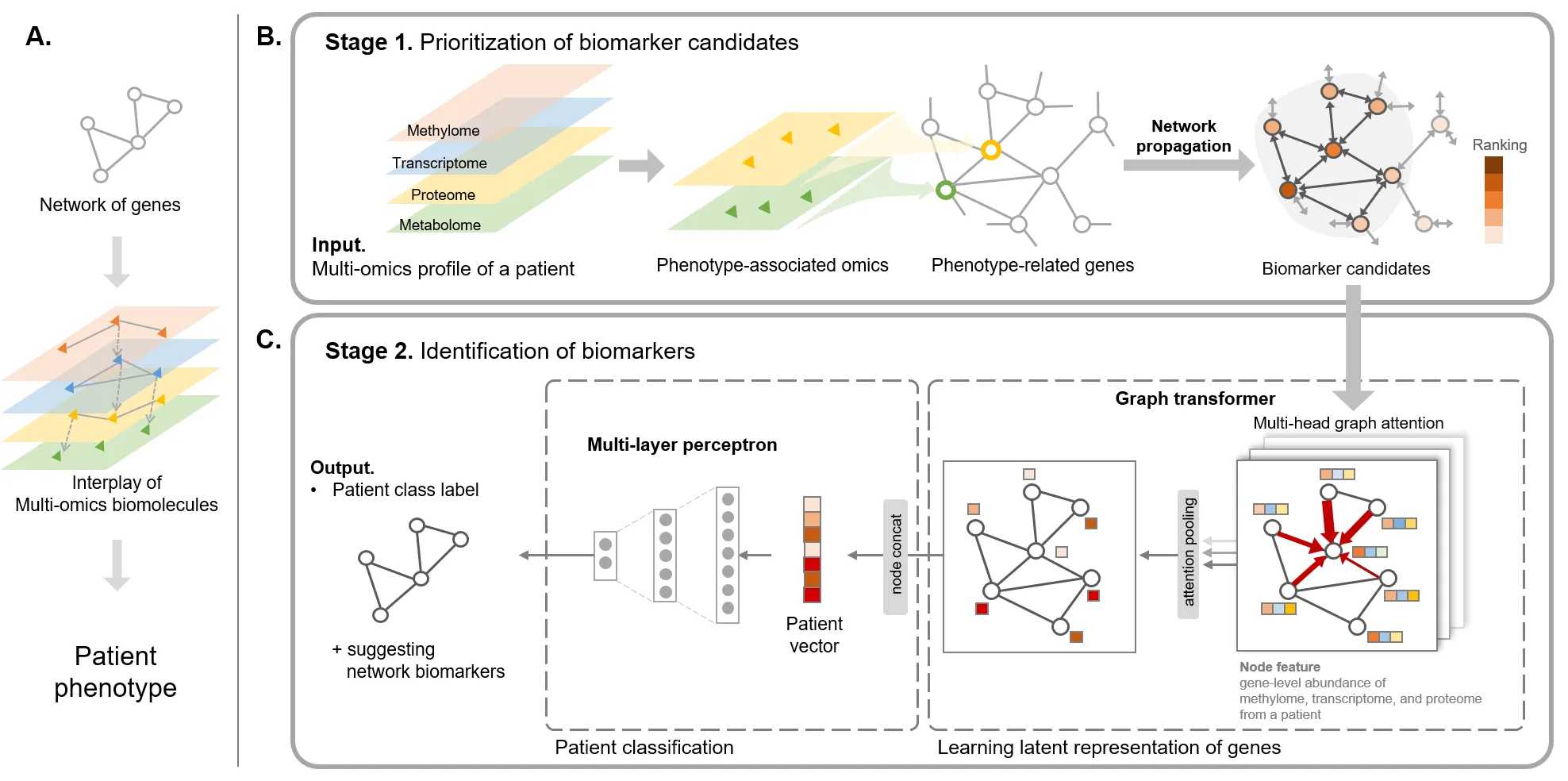
2026, ECCB (Under revision) “MIL2Het: Learning Patient Phenotypes from Single-Cell Heterogeneity with Multi-view Prior Knowledge-informed Graph Learning”
•
2024, BMC genomics (Under revision) “ALPACA: A Visual Data Mining System for Subcellular Location-specific Knowledge Mining from Multi-Omics Data in Cancer”
•
Co-lead 2025, BioData Mining “Identification of Severity Related Mutation Hotspots in SARS-CoV-2 Using a Density-Based Clustering Approach”
•
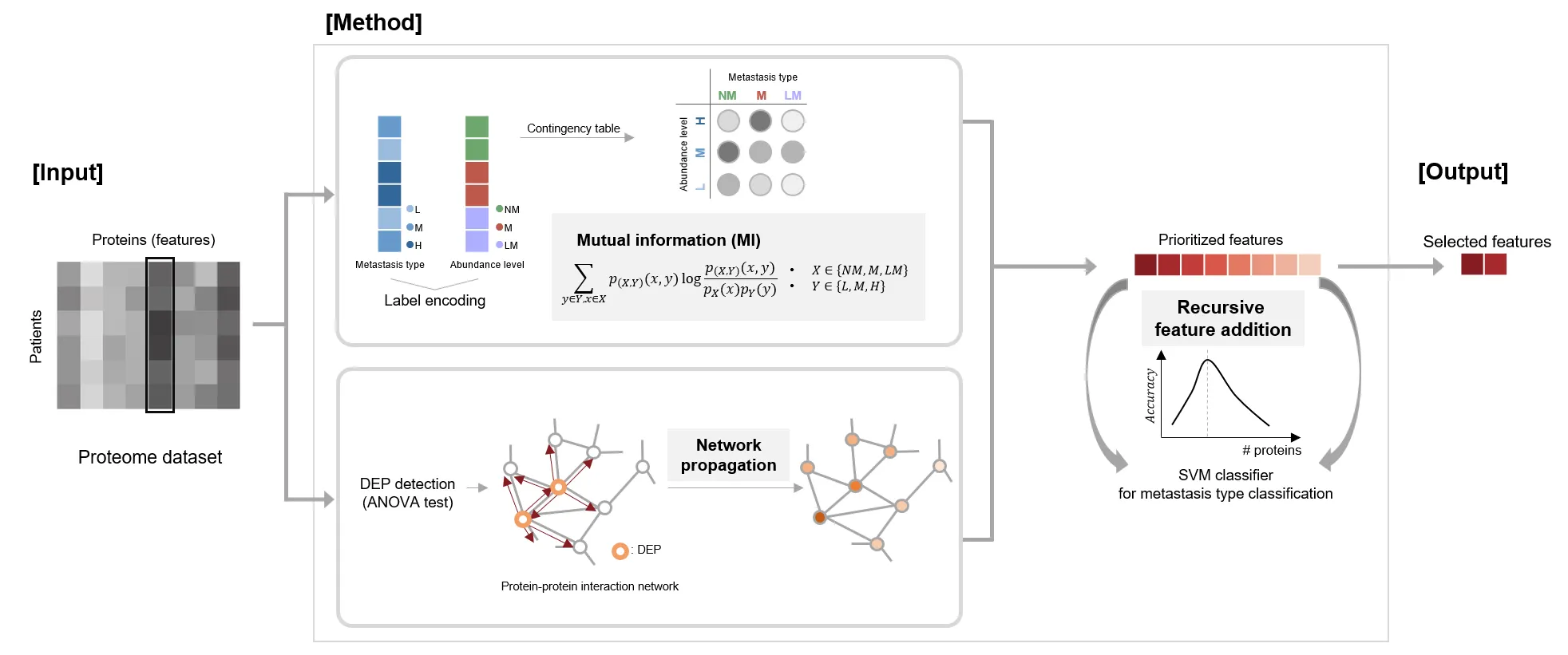
Co-lead 2024, Scientific Reports “Identification of VWA5A as a Novel Biomarker for Inhibiting Metastasis in Breast Cancer by Machine-learning Based Protein Prioritization”
•
2023, Proceedings of the 14th ACM International Conference on Bioinformatics, Computational Biology and Health Informatics “Deep learning-based survival prediction using DNA methylation-derived 3D genomic information”
Presented by Dabin Jeong at ACM-BCB 2023
•
2023, Journal of Korean Medical Science “Machine Learning-Based Proteomics Reveals Ferroptosis in COPD Patient-Derived Airway Epithelial Cells Upon Smoking Exposure”
•
2022, International Journal of Molecular Science “A Survey on Computational Methods for Investigation on ncRNA-Disease Association through the Mode of Action Perspective”
•
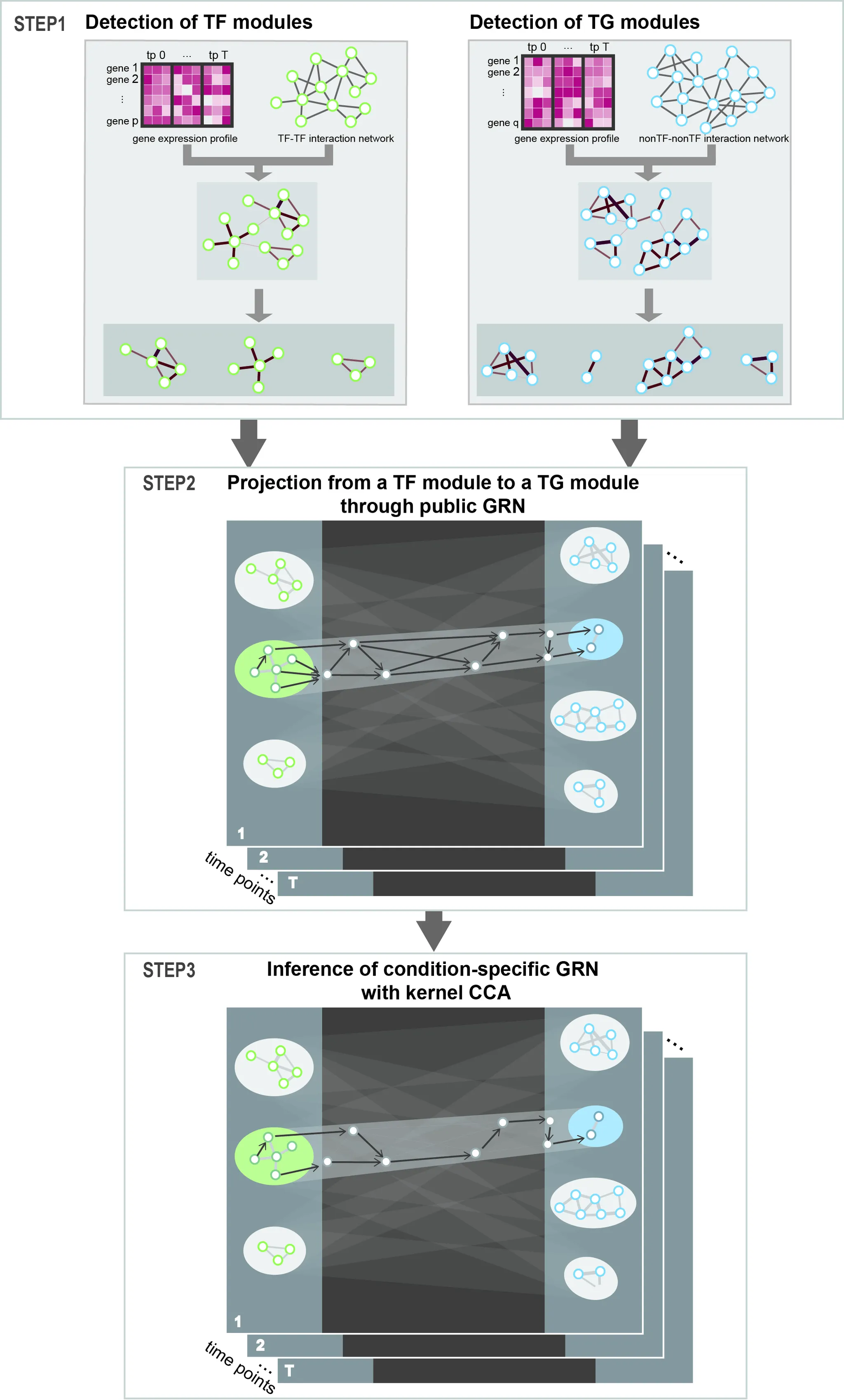
Lead 2021, Frontiers in Genetics “Construction of Condition-Specific Gene Regulatory Network using Kernel Canonical Correlation Analysis”
•
Co-lead 2020, Recent Advances in Biological network Analysis, “Network Propagation for the Analysis of Multi-omics Data”
•
Presented by Dabin Jeong at Genome Informatics Workshop (GIW) 2019
•
2019, Frontiers in Plant Science “PropaNet: Time-Varying Condition-Specific Transcriptional Network Construction by Network Propagation”
•
2018, Nucleic Acids Research “TRRUST v2: an expanded reference database of human and mouse transcriptional regulatory interactions”
cited 1,831 times (25.9.11)
Portfolio
Skills
•
Python
•
R
•
Docker
•
Nextflow
•
Snakemake
•
Git
•
Shell script